Biography
Contact details:
- +44 (0) 1603 450 036
Richard is Group Leader of the Technology Algorithms Group (Leggett Group).
His research interests include:
- Application of new sequencing technologies
- Real-time sequence analysis
- Sequencing for diagnostics and surveillance
- Classification and assembly of metagenomic samples
- In-field and in situ sequencing
- Development of new bioinformatics tools
- Detection of pathogens from the air using Air-seq
- Pathogen and AMR analysis of clinical metagenomic samples
He has been involved in the development of a number of bioinformatics tools, including:
Graduating in Physics, Richard spent 10 years working as a software engineer, before undertaking an MSc in Advanced Computing Science and a PhD in Computational Biology.
His PhD thesis was entitled, "Computational approaches for the analysis and modelling of filamentous growth and branching of Steptomyces coelicolor".
Following the PhD, and prior to joining EI, Richard was a postdoc at The Sainsbury Laboratory, Norwich, looking at novel methods for SNP detection in reference-free organisms.
Projects
Publications
Related reading.
Image

12 December 2022
Technology
FEATURE
| 10 min READ
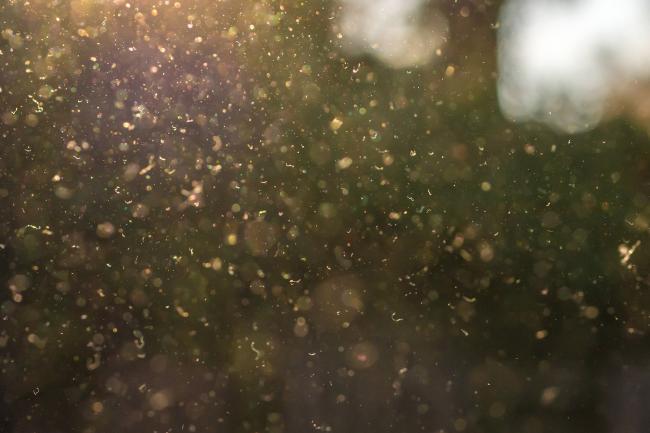
Sky’s the limit for new air sequencing technology
There’s something in the air. Literally.
Image

30 January 2019
Science
FEATURE
| 5 min READ
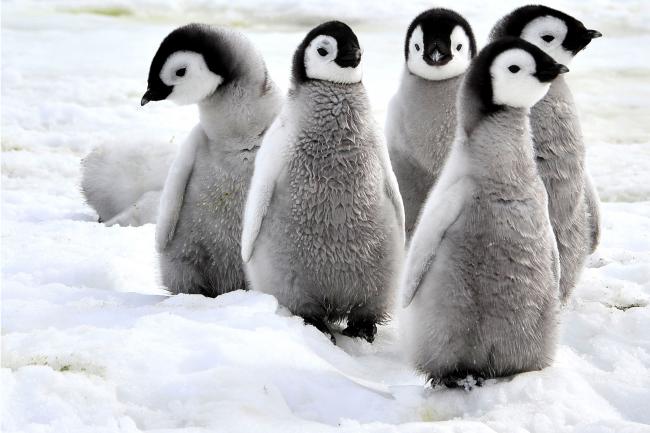
NanoOK of the North and South: A story of life in the Antarctic
Technology and Algorithms Group Leader Dr Richard Leggett tells us about his pioneering software tools NanoOK & NanoOK RT, which are making an impact in life sciences research.
Image

24 September 2018
Science
FEATURE
| 3 min READ
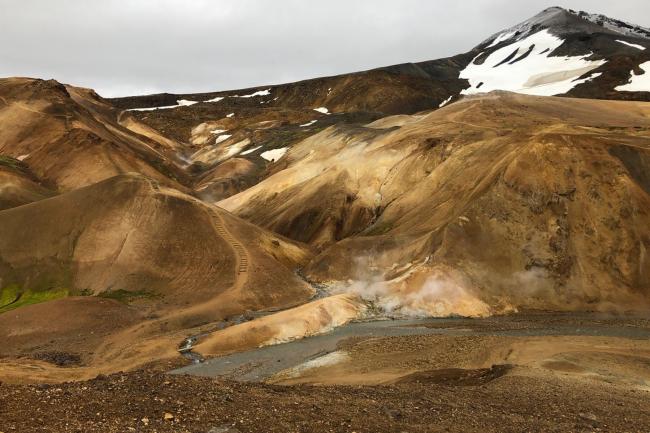
Extreme Environments: Genome sequencing & Space
Richard Leggett has recently returned from Iceland, where he’s been helping space scientists get to grips with genome sequencing in extreme environments.
Image

04 September 2020
Science
FEATURE
| 8 min READ
Metagenomics: exploring the unseen
Earlham Institute Scientists are at the forefront of one of biology’s modern frontiers: the untraversed depths of nature’s hidden microbial universe. From the bacteria living inside of us to the millions of single celled protists underpinning the health and prosperity of our ecosystems.
17 May 2021
Image

When one become two: separating DNA for more accurate nanopore analysis
A new software tool developed by Earlham Institute researchers will help bioinformaticians improve the quality and accuracy of their biological data, and avoid mis-assemblies.
16 December 2019
Image

Diagnosing infections earlier in preterm babies with real time genomic analysis
Scientists at the Norwich Research Park pioneer microbiome profiling method to speed up ID of infections and more effective treatments in preterm babies.
07 July 2020
Image

What's living in the River Yare?
Rivers are the lifeblood of civilisation. It's easy to see why so many of the world's major cities were built around waterways and their fertile, alluvial plains, and Norwich is no different.